...
| Note |
|---|
This is a new service and these are new directions, if something doesn't work or is missing from the documentation please contact me so I can fix it. |
...
- Go to https://processing.crbs.ucsd.edu/ and log in using your CRBS SSO (LDAP) credentials
- Click on Run a standardized process on a dataset
- Click on MRC operations
- Enter the full path to the stack you want part of
- If you have the MPID or Project number you can use the link at the top of the page to quickly find your stack. Once found you can copy the path to the clipboard or use the Get stack info button to take you back to the portal
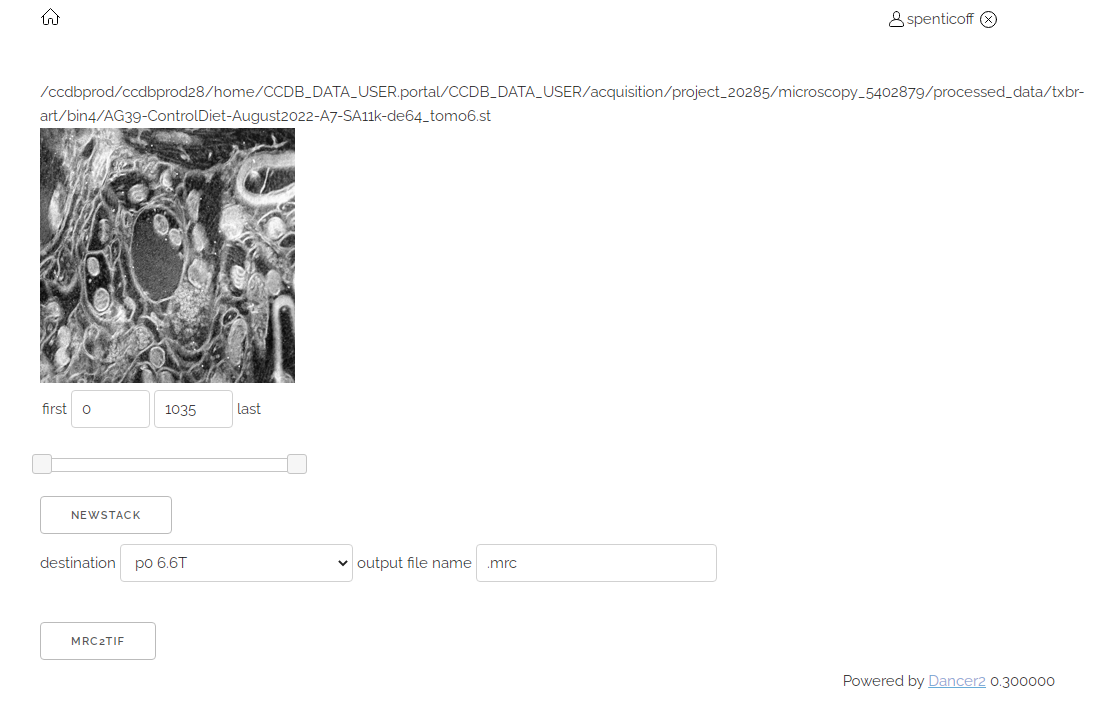
- It may take a few moments to get the information about the stack but you should eventually see the following
- You can use the slider or type in the first and last boxes to choose the subset of layers you want.
- Choose a destination
- p0-p5 are mounted on all of the linux workstations under /processing
- choose download temp_filesystem if you will be downloading the file for processing elsewhere, files that go here are automatically deleted when they are 5 days old
- Be sure to name your file
- Press the newstack button. You will be given some information and asked to confirm or cancel
- When you press confirm the job will be queued up and run and the file will appear under the filesystem you chose (this can take seconds to minutes)
...
?path=/ccdbprod/ccdbprod28/home/CCDB_DATA_USER.portal/CCDB_DATA_USER/acquisition/project_20285/microscopy_5402879/processed_data/txbr-art/bin4/AG39-ControlDiet-August2022-A7-SA11k-de64_tomo6.st
&first=0
&last=15
&filename=for_download.mrc
(be sure to include the credentials or you will get back a 403)
All 4 parameters are required, the file will always be in the download directory and the download URL for this example would be
...
| Page properties | ||
|---|---|---|
| ||
|